Ribosome
| Cell biology | |
|---|---|
| The animal cell | |
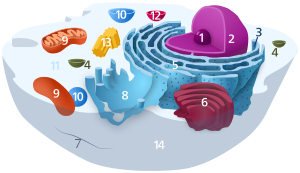 Components of a typical animal cell:
|

Figure 1: Ribosomes assemble polymeric protein molecules whose sequence is controlled by the sequence of messenger RNA molecules. This is required by all living cells and associated viruses.
The ribosome (/ˈraɪbəˌsoʊm, -boʊ-/[1]) is a complex molecular machine, found within all living cells, that serves as the site of biological protein synthesis (translation). Ribosomes link amino acids together in the order specified by messenger RNA (mRNA) molecules. Ribosomes consist of two major components: the small ribosomal subunits, which read the RNA, and the large subunits, which join amino acids to form a polypeptide chain. Each subunit consists of one or more ribosomal RNA (rRNA) molecules and a variety of ribosomal proteins (r-protein or rProtein[2][3][4]). The ribosomes and associated molecules are also known as the translational apparatus.
Contents
1 Overview
2 Discovery
3 Structure
3.1 Prokaryotic ribosomes
3.2 Eukaryotic ribosomes
3.3 Plastoribosomes and mitoribosomes
3.4 Making use of the differences
3.5 Common properties
3.6 High-resolution structure
4 Function
4.1 Translation
4.1.1 Cotranslational folding
4.2 Addition of translation-independent amino acids
5 Ribosome locations
5.1 Free ribosomes
5.2 Membrane-bound ribosomes
6 Biogenesis
7 Origin
8 Specialized ribosomes
8.1 Ribosomal proteins
8.2 Ribosome-associated factors
8.3 rRNA heterogeneity
8.4 rRNA modifications
9 See also
10 References
11 External links
Overview
The sequence of DNA, which encodes the sequence of the amino acids in a protein, is copied into a messenger RNA chain. It may be copied many times into RNA chains. Ribosomes can bind to a messenger RNA chain and use its sequence for determining the correct sequence of amino acids for generating a given protein. Amino acids are selected, collected, and carried to the ribosome by transfer RNA (tRNA) molecules, which enter one part of the ribosome and bind to the messenger RNA chain. It is during this binding that the correct translation of nucleic acid sequence to amino acid sequence occurs. For each coding triplet in the messenger RNA there is a distinct transfer RNA that matches and which carries the correct amino acid for that coding triplet. The attached amino acids are then linked together by another part of the ribosome. Once the protein is produced, it can then fold to produce a specific functional three-dimensional structure although during synthesis some proteins start folding into their correct form.
A ribosome is made from complexes of RNAs and proteins and is therefore a ribonucleoprotein. Each ribosome is divided into two subunits:
- a smaller subunit which binds to a larger subunit and the mRNA pattern, and
- a larger subunit which binds to the tRNA, the amino acids, and the smaller subunit.
When a ribosome finishes reading an mRNA molecule, these two subunits split apart. Ribosomes are ribozymes, because the catalytic peptidyl transferase activity that links amino acids together is performed by the ribosomal RNA. Ribosomes are often associated with the intracellular membranes that make up the rough endoplasmic reticulum.
Ribosomes from bacteria, archaea and eukaryotes in the three-domain system, resemble each other to a remarkable degree, evidence of a common origin. They differ in their size, sequence, structure, and the ratio of protein to RNA. The differences in structure allow some antibiotics to kill bacteria by inhibiting their ribosomes, while leaving human ribosomes unaffected. In bacteria and archaea, more than one ribosome may move along a single mRNA chain at one time, each "reading" its sequence and producing a corresponding protein molecule.
The mitochondrial ribosomes of eukaryotic cells, are produced from mitochondrial genes, and functionally resemble many features of those in bacteria, reflecting the likely evolutionary origin of mitochondria.[5][6]
Discovery
Ribosomes were first observed in the mid-1950s by Romanian-American cell biologist George Emil Palade, using an electron microscope, as dense particles or granules.[7] The term "ribosome" was proposed by scientist Richard B. Roberts in the end of 1950s:
.mw-parser-output .templatequoteoverflow:hidden;margin:1em 0;padding:0 40px.mw-parser-output .templatequote .templatequoteciteline-height:1.5em;text-align:left;padding-left:1.6em;margin-top:0
During the course of the symposium a semantic difficulty became apparent. To some of the participants, "microsomes" mean the ribonucleoprotein particles of the microsome fraction contaminated by other protein and lipid material; to others, the microsomes consist of protein and lipid contaminated by particles. The phrase "microsomal particles" does not seem adequate, and "ribonucleoprotein particles of the microsome fraction" is much too awkward. During the meeting, the word "ribosome" was suggested, which has a very satisfactory name and a pleasant sound. The present confusion would be eliminated if "ribosome" were adopted to designate ribonucleoprotein particles in sizes ranging from 35 to 100S.
— Albert, Microsomal Particles and Protein Synthesis[8]
Albert Claude, Christian de Duve, and George Emil Palade were jointly awarded the Nobel Prize in Physiology or Medicine, in 1974, for the discovery of the ribosome.[9] The Nobel Prize in Chemistry 2009 was awarded to Venkatraman Ramakrishnan, Thomas A. Steitz and Ada E. Yonath for determining the detailed structure and mechanism of the ribosome.[10]
Structure
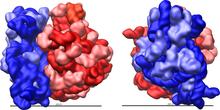
Figure 2: Large (red) and small (blue) subunit fit together.
The ribosome is a highly complex cellular machine. It is largely made up of specialized RNA known as ribosomal RNA (rRNA) as well as dozens of distinct proteins (the exact number varies slightly between species). The ribosomal proteins and rRNAs are arranged into two distinct ribosomal pieces of different size, known generally as the large and small subunit of the ribosome. Ribosomes consist of two subunits that fit together (Figure 2) and work as one to translate the mRNA into a polypeptide chain during protein synthesis (Figure 1). Because they are formed from two subunits of non-equal size, they are slightly longer in the axis than in diameter.
Prokaryotic ribosomes
Prokaryotic ribosomes are around 20 nm (200 Å) in diameter and are composed of 65% rRNA and 35% ribosomal proteins.[11] Eukaryotic ribosomes are between 25 and 30 nm (250–300 Å) in diameter with an rRNA-to-protein ratio that is close to 1.[12]Crystallographic work [13] has shown that there are no ribosomal proteins close to the reaction site for polypeptide synthesis. This suggests that the protein components of ribosomes do not directly participate in peptide bond formation catalysis, but rather that these proteins act as a scaffold that may enhance the ability of rRNA to synthesize protein (See: Ribozyme).

Figure 3: Atomic structure of the 30S subunit from Thermus thermophilus.[14] Proteins are shown in blue and the single RNA chain in brown.
The ribosomal subunits of prokaryotes and eukaryotes are quite similar.[15]
The unit of measurement used to describe the ribosomal subunits and the rRNA fragments is the Svedberg unit, a measure of the rate of sedimentation in centrifugation rather than size. This accounts for why fragment names do not add up: for example, prokaryotic 70S ribosomes are made of 50S and 30S subunits.
Prokaryotes have 70S ribosomes, each consisting of a small (30S) and a large (50S) subunit. Their small subunit has a 16S RNA subunit (consisting of 1540 nucleotides) bound to 21 proteins. The large subunit is composed of a 5S RNA subunit (120 nucleotides), a 23S RNA subunit (2900 nucleotides) and 31 proteins.[15]
prokaryotic ribosomes (E. coli)[16] ribosome subunit rRNAs r-proteins 70S 50S 23S (2904 nt) 31 5S (120 nt) 30S 16S (1542 nt) 21
Affinity label for the tRNA binding sites on the E. coli ribosome allowed the identification of A and P site proteins most likely associated with the peptidyltransferase activity; labelled proteins are L27, L14, L15, L16, L2; at least L27 is located at the donor site, as shown by E. Collatz and A.P. Czernilofsky.[17][18] Additional research has demonstrated that the S1 and S21 proteins, in association with the 3′-end of 16S ribosomal RNA, are involved in the initiation of translation.[19]
Eukaryotic ribosomes
Eukaryotes have 80S ribosomes located in their cytosol, each consisting of a small (40S) and large (60S) subunit. Their 40S subunit has an 18S RNA (1900 nucleotides) and 33 proteins.[20][21] The large subunit is composed of a 5S RNA (120 nucleotides), 28S RNA (4700 nucleotides), a 5.8S RNA (160 nucleotides) subunits and 46 proteins.[15][20][22]
eukaryotic cytosolic ribosomes (R. norvegicus)[23] ribosome subunit rRNAs r-proteins 80S 60S 28S (4718 nt) 49 5.8S (160 nt) 5S (120 nt) 40S 18S (1874 nt) 33
During 1977, Czernilofsky published research that used affinity labeling to identify tRNA-binding sites on rat liver ribosomes. Several proteins, including L32/33, L36, L21, L23, L28/29 and L13 were implicated as being at or near the peptidyl transferase center.[24]
Plastoribosomes and mitoribosomes
In eukaryotes there are ribosomes found in mitochondria (sometimes called mitoribosomes) and in plastids such as chloroplasts (also called plastoribosomes). They also consist of large and small subunits bound together with proteins into one 70S particle.[15] These organelles are believed to be descendants of bacteria (see Endosymbiotic theory) and, as such, their ribosomes are similar to those of bacteria.[15] Of the two, chloroplastic ribosomes are closer to bacterial ones than mitochrondrial ones are. Many pieces of ribosomal RNA in the mitochrondria are shortened, and in the case of 5S rRNA, replaced by other structures in animals and fungi.[25] In particular, Leishmania tarentolae has a minimalized set of mitochondrial rRNA.[26]
Note that komplex plastids may contain a nucleomorph which resembles a small eukaryotic nucleus. Subsequently, in the compartiment nearby the nucleomorph, there are eukaryotic 80S ribosomes to be found, too.
Making use of the differences
The differences between the bacterial and eukaryotic ribosomes are exploited by pharmaceutical chemists to create antibiotics that can destroy a bacterial infection without harming the cells of the infected person. Due to the differences in their structures, the bacterial 70S ribosomes are vulnerable to these antibiotics while the eukaryotic 80S ribosomes are not.[27] Even though mitochondria possess ribosomes similar to the bacterial ones, mitochondria are not affected by these antibiotics because they are surrounded by a double membrane that does not easily admit these antibiotics into the organelle.[28] The same cannot be said of chloroplasts, where antibiotic resistance in ribosomal proteins is a trait to be introduced as a marker in genetic engineering.[29]
Common properties
The various ribosomes share a core structure, which is quite similar despite the large differences in size. Much of the RNA is highly organized into various tertiary structural motifs, for example pseudoknots that exhibit coaxial stacking. The extra RNA in the larger ribosomes is in several long continuous insertions, such that they form loops out of the core structure without disrupting or changing it.[15] All of the catalytic activity of the ribosome is carried out by the RNA; the proteins reside on the surface and seem to stabilize the structure.[15]
High-resolution structure

Figure 4: Atomic structure of the 50S subunit from Haloarcula marismortui. Proteins are shown in blue and the two RNA chains in brown and yellow.[30] The small patch of green in the center of the subunit is the active site.
The general molecular structure of the ribosome has been known since the early 1970s. In the early 2000s, the structure has been achieved at high resolutions, of the order of a few ångströms.
The first papers giving the structure of the ribosome at atomic resolution were published almost simultaneously in late 2000. The 50S (large prokaryotic) subunit was determined from the archaeon Haloarcula marismortui[30] and the bacterium Deinococcus radiodurans,[31] and the structure of the 30S subunit was determined from Thermus thermophilus.[14] These structural studies were awarded the Nobel Prize in Chemistry in 2009. In May 2001 these coordinates were used to reconstruct the entire T. thermophilus 70S particle at 5.5 Å resolution.[32]
Two papers were published in November 2005 with structures of the Escherichia coli 70S ribosome. The structures of a vacant ribosome were determined at 3.5 Å resolution using X-ray crystallography.[33] Then, two weeks later, a structure based on cryo-electron microscopy was published,[34] which depicts the ribosome at 11–15 Å resolution in the act of passing a newly synthesized protein strand into the protein-conducting channel.
The first atomic structures of the ribosome complexed with tRNA and mRNA molecules were solved by using X-ray crystallography by two groups independently, at 2.8 Å[35] and at 3.7 Å.[36] These structures allow one to see the details of interactions of the Thermus thermophilus ribosome with mRNA and with tRNAs bound at classical ribosomal sites. Interactions of the ribosome with long mRNAs containing Shine-Dalgarno sequences were visualized soon after that at 4.5–5.5 Å resolution.[37]
In 2011, the first complete atomic structure of the eukaryotic 80S ribosome from the yeast Saccharomyces cerevisiae was obtained by crystallography.[20] The model reveals the architecture of eukaryote-specific elements and their interaction with the universally conserved core. At the same time, the complete model of a eukaryotic 40S ribosomal structure in Tetrahymena thermophila was published and described the structure of the 40S subunit, as well as much about the 40S subunit's interaction with eIF1 during translation initiation.[21] Similarly, the eukaryotic 60S subunit structure was also determined from Tetrahymena thermophila in complex with eIF6.[22]
Function
Ribosomes are minute particles consisting of RNA and associated proteins that function to synthesize proteins. Proteins are needed for many cellular functions such as repairing damage or directing chemical processes. Ribosomes can be found floating within the cytoplasm or attached to the endoplasmic reticulum.
Ribosomes act as catalysts in two extremely important biological processes called peptidyl transfer and peptidyl hydrolysis.[38] The "PT center is responsible for producing protein bonds during protein elongation".[38]
Translation
Ribosomes are the workplaces of protein biosynthesis, the process of translating mRNA into protein. The mRNA comprises a series of codons that dictate to the ribosome the sequence of the amino acids needed to make the protein. Using the mRNA as a template, the ribosome traverses each codon (3 nucleotides) of the mRNA, pairing it with the appropriate amino acid provided by an aminoacyl-tRNA. Aminoacyl-tRNA contains a complementary anticodon on one end and the appropriate amino acid on the other. For fast and accurate recognition of the appropriate tRNA, the ribosome utilizes large conformational changes (conformational proofreading)
.[39]
The small ribosomal subunit, typically bound to an aminoacyl-tRNA containing the amino acid methionine, binds to an AUG codon on the mRNA and recruits the large ribosomal subunit. The ribosome contains three RNA binding sites, designated A, P and E. The A-site binds an aminoacyl-tRNA;[40] the P-site binds a peptidyl-tRNA (a tRNA bound to the peptide being synthesized); and the E-site (exit) binds a free tRNA before it exits the ribosome. Protein synthesis begins at a start codon AUG near the 5' end of the mRNA. mRNA binds to the P site of the ribosome first. The ribosome is able to identify the start codon by use of the Shine-Dalgarno sequence of the mRNA in prokaryotes and Kozak box in eukaryotes.
Although catalysis of the peptide bond involves the C2 hydroxyl of RNA's P-site adenosine in a proton shuttle mechanism, other steps in protein synthesis (such as translocation) are caused by changes in protein conformations. Since their catalytic core is made of RNA, ribosomes are classified as "ribozymes,"[41] and it is thought that they might be remnants of the RNA world.[42]

Figure 5: Translation of mRNA (1) by a ribosome (2)(shown as small and large subunits) into a polypeptide chain (3). The ribosome begins at the start codon of RNA (AUG) and ends at the stop codon (UAG).
In Figure 5, both ribosomal subunits (small and large) assemble at the start codon (towards the 5' end of the RNA). The ribosome uses RNA that matches the current codon (triplet) on the mRNA to append an amino acid to the polypeptide chain. This is done for each triplet on the RNA, while the ribosome moves towards the 3' end of the mRNA. Usually in bacterial cells, several ribosomes are working parallel on a single RNA, forming what is called a polyribosome or polysome.
Cotranslational folding
The ribosome is known to actively participate in the protein folding.[43] The structures obtained in this way are usually identical to the ones obtained during protein chemical refolding, however, the pathways leading to the final product may be different.[44] In some cases, the ribosome is crucial in obtaining the functional protein form. For example, one of the possible mechanisms of folding of the deeply knotted proteins relies on the ribosome pushing the chain through the attached loop.[45]
Addition of translation-independent amino acids
Presence of a ribosome quality control protein Rqc2 is associated with mRNA-independent protein elongation.[46][47] This elongation is a result of ribosomal addition (via tRNAs brought by Rqc2) of CAT tails: ribosomes extend the C-terminus of a stalled protein with random, translation-independent sequences of alanines and threonines.[48][49]
Würzburg University and Max Planck Institute researches, the results of which were published in Cell Reports and The EMBO magazines in September 2016, have shown that ribosomes have the role of being "a quality control point". Professor Utz Fischer from the University of Würzburg has been researching the assembly of proteins called "macromolecular machines" in the cell for years. He describes this assembly process as LEGO blocks: "Think of it as LEGO bricks at the molecular level: One brick is attached to the next until the product is finished. If only one defective or wrong brick is used, the entire building may be compromised as a result."[50][51][52]
Ribosome locations
Ribosomes are classified as being either "free" or "membrane-bound".

Figure 6: A ribosome translating a protein that is secreted into the endoplasmic reticulum.
Free and membrane-bound ribosomes differ only in their spatial distribution; they are identical in structure. Whether the ribosome exists in a free or membrane-bound state depends on the presence of an ER-targeting signal sequence on the protein being synthesized, so an individual ribosome might be membrane-bound when it is making one protein, but free in the cytosol when it makes another protein.
Ribosomes are sometimes referred to as organelles, but the use of the term organelle is often restricted to describing sub-cellular components that include a phospholipid membrane, which ribosomes, being entirely particulate, do not. For this reason, ribosomes may sometimes be described as "non-membranous organelles".
Free ribosomes
Free ribosomes can move about anywhere in the cytosol, but are excluded from the cell nucleus and other organelles. Proteins that are formed from free ribosomes are released into the cytosol and used within the cell. Since the cytosol contains high concentrations of glutathione and is, therefore, a reducing environment, proteins containing disulfide bonds, which are formed from oxidized cysteine residues, cannot be produced within it.
Membrane-bound ribosomes
When a ribosome begins to synthesize proteins that are needed in some organelles, the ribosome making this protein can become "membrane-bound". In eukaryotic cells this happens in a region of the endoplasmic reticulum (ER) called the "rough ER". The newly produced polypeptide chains are inserted directly into the ER by the ribosome undertaking vectorial synthesis and are then transported to their destinations, through the secretory pathway. Bound ribosomes usually produce proteins that are used within the plasma membrane or are expelled from the cell via exocytosis.[53]
Biogenesis
In bacterial cells, ribosomes are synthesized in the cytoplasm through the transcription of multiple ribosome gene operons. In eukaryotes, the process takes place both in the cell cytoplasm and in the nucleolus, which is a region within the cell nucleus. The assembly process involves the coordinated function of over 200 proteins in the synthesis and processing of the four rRNAs, as well as assembly of those rRNAs with the ribosomal proteins.
Origin
The ribosome may have first originated in an RNA world, appearing as a self-replicating complex that only later evolved the ability to synthesize proteins when amino acids began to appear.[54] Studies suggest that ancient ribosomes constructed solely of rRNA could have developed the ability to synthesize peptide bonds.[55][56][57] In addition, evidence strongly points to ancient ribosomes as self-replicating complexes, where the rRNA in the ribosomes had informational, structural, and catalytic purposes because it could have coded for tRNAs and proteins needed for ribosomal self-replication.[58] Hypothetical cellular organisms with self-replicating RNA but without DNA are called ribocytes (or ribocells).[59][60]
As amino acids gradually appeared in the RNA world under prebiotic conditions,[61][62] their interactions with catalytic RNA would increase both the range and efficiency of function of catalytic RNA molecules.[54] Thus, the driving force for the evolution of the ribosome from an ancient self-replicating machine into its current form as a translational machine may have been the selective pressure to incorporate proteins into the ribosome’s self-replicating mechanisms, so as to increase its capacity for self-replication.[58]
Specialized ribosomes
Many textbooks suggest that there are only two kinds of ribosomes, namely prokaryotic and eukaryotic ribosomes. However, ribosomes are surprisingly heterogeneous with different compositions in different species. Heterogeneous ribosomes have different structures and thus different activities compared to typical ribosomes in major model organisms.

Figure 7: The location of specialized ribosomal proteins (RPs) on the yeast ribosome on the small ribosomal unit (SSU)and large ribosomal subunit (LSU). Highlighted in green are the RPs on the SSU that increase translation of wild type Renilla luciferase (REN) gene and FF (Firefly) luciferase. Highlighted in green are the RPs on LSU that increase the expression of LAMB3 reporter with premature termination codons (PTC).[63]
Heterogeneity in ribosome composition has been proposed to be involved in translational control of protein synthesis.[64] Vincent Mauro and Gerald Edelman proposed the ribosome filter hypothesis to explain the regulatory functions of ribosomes. Emerging evidence has shown that specialized ribosomes specific to different cell populations can affect how genes are translated.[65] Some ribosomal proteins exchange from the assembled complex with cytosolic copies [66] suggesting that the structure of the in vivo ribosome can be modified without synthesizing an entire new ribosome.
Ribosomal proteins
A group of highly acidic ribosomal proteins (RPs), also known as P proteins, are known to be present on the 60S subunit in multiple copies in the ribosomes stalk and P proteins mediate selective translation.[67] These P proteins can be found in yeast and mammalian cells. If P proteins are not present in yeast this can cause the yeast to have a cold-sensitive phenotype.[68] If P proteins are not present in human cells, this could cause autophagy induction.[69]
Certain ribosomal proteins are absolutely critical while others are not. For instance, Rpl28 and Rpl5 mutant flies are alive but have abnormally large wings.[70] Rpl38 also appears to be critical in mammals only under very specific conditions: in mice Rpl38 is required for the translation of a subset of Hox mRNA and a mutation of Rpl38 lead to a homeotic transformation with a short tail.[71]
Other ways heterogeneity can arise from post-translational modifications to RPs include acetylation and methylation in species like yeast,[72]Arabidopsis,[73] and human cells.[74][75][76] But there is no overall change in protein synthesis.
Modifications to core ribosomal proteins (RPs) can also give rise to the formation of heterogeneous ribosomes. For instance, in yeast, Rpl28 ubiquitination levels vary with the cell cycle. Ribosomes with the polyubiquitinated Rpl28 carry out protein synthesis at a higher rate in vitro compared with ribosomes with monoubiquitinated Rpl28.[77]

Figure 8: Types of ribosome heterogeneity and its components.[78]
Ribosome-associated factors
Heterogeneous ribosomes can also arise from other factors binding to the ribosome surface. For example, ribosome-associated factor (RACK1) is so tightly associated with the ribosome that its binding is resistant to high-salt washes in vitro. RACK1 is required for efficient translation of mRNA with short open reading frames.[79]
rRNA heterogeneity
Viral IRES (internal ribosome entry site) translations can also be mediated by specialized ribosomes. Specifically, 40S ribosomal units without RPS25 in yeast and mammalian cells are unable to begin translation from two viral IRESes, namely the hepatitis C virus IRES and the cricket paralysis virus (CrPV) intergenic region. The CrPV only needs RPS25 to begin translation and mediate ribosome recruitment. Usually if RPS25 is not present in a certain IRES, imitation from these IRESes can be defective.[80][81][82]
rRNA modifications
Heterogeneity of ribosomal RNA modifications plays an important role in structural maintenance and/or function, such as modulating translation and most mRNA modifications are found in highly conserved regions.[83][84] The most common rRNA modifications are pseudouridylation and 2’-O methylation of ribose.[67]
See also
- Aminoglycosides
- Biological machines
- Eukaryotic translation
- Posttranslational modification
- Prokaryotic translation
- Protein dynamics
- RNA tertiary structure
- Translation (genetics)
- Wobble base pair
References
^ Jones, Daniel (2003) [1917], Peter Roach, James Hartmann and Jane Setter, eds., English Pronouncing Dictionary, Cambridge: Cambridge University Press, ISBN 978-3-12-539683-8CS1 maint: Uses editors parameter (link).mw-parser-output cite.citationfont-style:inherit.mw-parser-output .citation qquotes:"""""""'""'".mw-parser-output .citation .cs1-lock-free abackground:url("//upload.wikimedia.org/wikipedia/commons/thumb/6/65/Lock-green.svg/9px-Lock-green.svg.png")no-repeat;background-position:right .1em center.mw-parser-output .citation .cs1-lock-limited a,.mw-parser-output .citation .cs1-lock-registration abackground:url("//upload.wikimedia.org/wikipedia/commons/thumb/d/d6/Lock-gray-alt-2.svg/9px-Lock-gray-alt-2.svg.png")no-repeat;background-position:right .1em center.mw-parser-output .citation .cs1-lock-subscription abackground:url("//upload.wikimedia.org/wikipedia/commons/thumb/a/aa/Lock-red-alt-2.svg/9px-Lock-red-alt-2.svg.png")no-repeat;background-position:right .1em center.mw-parser-output .cs1-subscription,.mw-parser-output .cs1-registrationcolor:#555.mw-parser-output .cs1-subscription span,.mw-parser-output .cs1-registration spanborder-bottom:1px dotted;cursor:help.mw-parser-output .cs1-ws-icon abackground:url("//upload.wikimedia.org/wikipedia/commons/thumb/4/4c/Wikisource-logo.svg/12px-Wikisource-logo.svg.png")no-repeat;background-position:right .1em center.mw-parser-output code.cs1-codecolor:inherit;background:inherit;border:inherit;padding:inherit.mw-parser-output .cs1-hidden-errordisplay:none;font-size:100%.mw-parser-output .cs1-visible-errorfont-size:100%.mw-parser-output .cs1-maintdisplay:none;color:#33aa33;margin-left:0.3em.mw-parser-output .cs1-subscription,.mw-parser-output .cs1-registration,.mw-parser-output .cs1-formatfont-size:95%.mw-parser-output .cs1-kern-left,.mw-parser-output .cs1-kern-wl-leftpadding-left:0.2em.mw-parser-output .cs1-kern-right,.mw-parser-output .cs1-kern-wl-rightpadding-right:0.2em
^ Salini Konikkat: Dynamic Remodeling Events Drive the Removal of the ITS2 Spacer Sequence During Assembly of 60S Ribosomal Subunits in S. cerevisiae. Archived 2017-08-03 at the Wayback Machine Carnegie Mellon University Dissertations, February 2016.
^ Elmar W. Weiler, Lutz Nover (2008), Allgemeine und molekulare Botanik (in German), Stuttgart: Georg Thieme Verlag, p. 532, ISBN 978-3-13-152791-2
^ Jesus de la Cruz, Katrin Karbstein, John L. Woolford, Jr. (2015), "Functions of Ribosomal Proteins in Assembly of Eukaryotic Ribosomes In Vivo", Annual review of biochemistry (in German), 84, pp. 93–129, doi:10.1146/annurev-biochem-060614-033917, PMC 4772166, PMID 25706898CS1 maint: Multiple names: authors list (link)
^ Benne R, Sloof P (1987). "Evolution of the mitochondrial protein synthetic machinery". BioSystems. 21 (1): 51–68. doi:10.1016/0303-2647(87)90006-2. PMID 2446672.
^ "Ribosomes". Archived from the original on 2009-03-20. Retrieved 2011-04-28.
^ Palade, G.E. (January 1955). "A small particulate component of the cytoplasm". J Biophys Biochem Cytol. 1 (1): 59–68. doi:10.1083/jcb.1.1.59. PMC 2223592. PMID 14381428.
^ Roberts, R. B., editor. (1958) "Introduction" in Microsomal Particles and Protein Synthesis. New York: Pergamon Press, Inc.
^ "The Nobel Prize in Physiology or Medicine 1974". Nobelprize.org. The Nobel Foundation. Archived from the original on 26 January 2013. Retrieved 10 December 2012.
^ "2009 Nobel Prize in Chemistry". The Nobel Foundation. Archived from the original on 8 April 2012. Retrieved 10 December 2012.
^ Kurland, C.G. (1960). "Molecular characterization of ribonucleic acid from Escherichia coli ribosomes". Journal of Molecular Biology. 2 (2): 83–91. doi:10.1016/s0022-2836(60)80029-0.
^ Wilson, Daniel N.; Doudna Cate, Jamie H. (May 2012). "The Structure and Function of the Eukaryotic Ribosome". Cold Spring Harbor Perspectives in Biology. 4 (5): a011536. doi:10.1101/cshperspect.a011536. ISSN 1943-0264. PMC 3331703. PMID 22550233.
^ Nissen, P.; Hansen, J.; Ban, N.; Moore, P.B.; Steitz, T.A. (2000-08-11). "The structural basis of ribosome activity in peptide bond synthesis". Science. 289 (5481): 920–30. doi:10.1126/science.289.5481.920. PMID 10937990.
^ ab Wimberly BT, Brodersen DE, Clemons WM Jr, Morgan-Warren RJ, Carter AP, Vonrhein C, Hartsch T, Ramakrishnan V (September 2000). "Structure of the 30S ribosomal subunit". Nature. 407 (6802): 327–39. doi:10.1038/35030006. PMID 11014182.
^ abcdefg Alberts, Bruce; et al. (2002). The Molecular Biology of the Cell (4th ed.). Garland Science. p. 342. ISBN 978-0-8153-3218-3.
^ Reginald Garrett and Charles M. Grisham: Biochemistry. (International Student Edition). Cengage Learning Services; 4th edition 2009;
ISBN 978-0-495-11464-2; p. 962.
^ Czernilofsky, A; Küchler, E; Stöffler, G.; Czernilofsky, P. (1976). "SITE OF REACTION ON RIBOSOMAL-PROTEIN L27 WITH AN AFFINITY LABEL DERIVATIVE OF TRANSFER-RNA-F(MET)". FEBS Letters. 63 (2): 283–286. doi:10.1016/0014-5793(76)80112-3. PMID 770196.
^ Czernilofsky, A; Collatz, E; Stöffler, G; Küchler, E (1974). "PROTEINS AT TRANSFER-RNA BINDING-SITES OF ESCHERICHIA-COLI RIBOSOMES". Proceedings of the National Academy of Sciences of the United States of America. 71 (1): 230–234. doi:10.1073/pnas.71.1.230. PMC 387971. PMID 4589893.
^ Czernilofsky, A; Kurland, C.G.; Stöffler, G. (1975). "30S ribosomal proteins associated with 3′-terminus of 16S RNA". FEBS Letters. 58 (1): 281–284. doi:10.1016/0014-5793(75)80279-1. PMID 1225593.
^ abc Ben-Shem A, Garreau de Loubresse N, Melnikov S, Jenner L, Yusupova G, Yusupov M (February 2011). "The structure of the eukaryotic ribosome at 3.0 Å resolution". Science. 334 (6062): 1524–1529. doi:10.1126/science.1212642. PMID 22096102.
^ ab Rabl, Leibundgut, Ataide, Haag, Ban (February 2010). "Crystal Structure of the Eukaryotic 40S Ribosomal Subunit in Complex with Initiation Factor 1". Science. 331 (6018): 730–736. doi:10.1126/science.1198308. hdl:20.500.11850/153130. PMID 21205638.CS1 maint: Multiple names: authors list (link)
^ ab Klinge, Voigts-Hoffmann, Leibundgut, Arpagaus, Ban (November 2011). "Crystal Structure of the Eukaryotic 60S Ribosomal Subunit in Complex with Initiation Factor 6". Science. 334 (6058): 941–948. doi:10.1126/science.1211204. PMID 22052974.CS1 maint: Multiple names: authors list (link)
^ Reginald Garrett and Charles M. Grisham: Biochemistry. (International Student Edition). Cengage Learning Services; 4th edition 2009;
ISBN 978-0-495-11464-2; p. 965.
^ Czernilofsky, A; Collatz, Ekkehard; Gressner, Axel M.; Wool, Ira G.; Küchler, Ernst (1977). "IDENTIFICATION OF TRNA-BINDING SITES ON RAT-LIVER RIBOSOMES BY AFFINITY LABELING". Molecular and General Genetics. 153 (3): 231–235. doi:10.1007/BF00431588.
^ Agrawal, RK; Sharma, MR (December 2012). "Structural aspects of mitochondrial translational apparatus". Current opinion in structural biology. 22 (6): 797–803. doi:10.1016/j.sbi.2012.08.003. PMC 3513651. PMID 22959417.
^ Sharma, MR; Booth, TM; Simpson, L; Maslov, DA; Agrawal, RK (16 June 2009). "Structure of a mitochondrial ribosome with minimal RNA". Proceedings of the National Academy of Sciences of the United States of America. 106 (24): 9637–42. doi:10.1073/pnas.0901631106. PMC 2700991. PMID 19497863.
^ Recht MI, Douthwaite S, Puglisi JD (1999). "Basis for bacterial specificity of action of aminoglycoside antibiotics". EMBO J. 18 (11): 3133–8. doi:10.1093/emboj/18.11.3133. PMC 1171394. PMID 10357824.
^ O'Brien, T.W. (1971). "The General Occurrence of 55S Ribosomes in Mammalian Liver Mitochondria". J. Biol. Chem. 245: 3409.
^ Newman, SM; Boynton, JE; Gillham, NW; Randolph-Anderson, BL; Johnson, AM; Harris, EH (December 1990). "Transformation of chloroplast ribosomal RNA genes in Chlamydomonas: molecular and genetic characterization of integration events". Genetics. 126 (4): 875–88. PMID 1981764.
^ ab Ban N, Nissen P, Hansen J, Moore P, Steitz T (2000). "The complete atomic structure of the large ribosomal subunit at 2.4 Å resolution". Science. 289 (5481): 905–20. CiteSeerX 10.1.1.58.2271. doi:10.1126/science.289.5481.905. PMID 10937989.
^ Schluenzen F, Tocilj A, Zarivach R, Harms J, Gluehmann M, Janell D, Bashan A, Bartels H, Agmon I, Franceschi F, Yonath A (September 1, 2000). "Structure of functionally activated small ribosomal subunit at 3.3 Å resolution". Cell. 102 (5): 615–23. doi:10.1016/S0092-8674(00)00084-2. PMID 11007480.
^ Yusupov MM, Yusupova GZ, Baucom A, et al. (May 2001). "Crystal structure of the ribosome at 5.5 Å resolution". Science. 292 (5518): 883–96. doi:10.1126/science.1060089. PMID 11283358.
^ Schuwirth BS, Borovinskaya MA, Hau CW, et al. (November 2005). "Structures of the bacterial ribosome at 3.5 Å resolution". Science. 310 (5749): 827–34. doi:10.1126/science.1117230. PMID 16272117.
^ Mitra K, Schaffitzel C, Shaikh T, et al. (November 2005). "Structure of the E. coli protein-conducting channel bound to a translating ribosome". Nature. 438 (7066): 318–24. doi:10.1038/nature04133. PMC 1351281. PMID 16292303.
^ Selmer M, Dunham CM, Murphy FV, et al. (September 2006). "Structure of the 70S ribosome complexed with mRNA and tRNA". Science. 313 (5795): 1935–42. doi:10.1126/science.1131127. PMID 16959973.
^ Korostelev A, Trakhanov S, Laurberg M, Noller HF (September 2006). "Crystal structure of a 70S ribosome-tRNA complex reveals functional interactions and rearrangements". Cell. 126 (6): 1065–77. doi:10.1016/j.cell.2006.08.032. PMID 16962654.
^ Yusupova G, Jenner L, Rees B, Moras D, Yusupov M (November 2006). "Structural basis for messenger RNA movement on the ribosome". Nature. 444 (7117): 391–4. doi:10.1038/nature05281. PMID 17051149.
^ ab "Specialized Internal Structures of Prokaryotes | Boundless Microbiology". courses.lumenlearning.com. Retrieved 2018-09-27.
^ Savir, Y; Tlusty, T (Apr 11, 2013). "The ribosome as an optimal decoder: a lesson in molecular recognition". Cell. 153 (2): 471–9. doi:10.1016/j.cell.2013.03.032. PMID 23582332.
^ Konevega, AL; Soboleva, NG; Makhno, VI; Semenkov, YP; Wintermeyer, W; Rodnina, MV; Katunin, VI (Jan 2004). "Purine bases at position 37 of tRNA stabilize codon-anticodon interaction in the ribosomal A site by stacking and Mg2+-dependent interactions". RNA. 10 (1): 90–101. doi:10.1261/rna.5142404. PMC 1370521. PMID 14681588.
^ Rodnina, M.V.; Beringer, M.; Wintermeyer, W. (January 2007). "How ribosomes make peptide bonds". Trends Biochem. Sci. 32 (1): 20–6. doi:10.1016/j.tibs.2006.11.007. PMID 17157507.
^ Cech, T. (August 11, 2000). "Structural biology. The Ribosome Is a Ribozyme". Science. 289 (5481): 878–9. doi:10.1126/science.289.5481.878. PMID 10960319.
^ Fedorov, Alexey N.; Baldwin, Thomas O. (1997-12-26). "Cotranslational Protein Folding". Journal of Biological Chemistry. 272 (52): 32715–32718. doi:10.1074/jbc.272.52.32715. ISSN 0021-9258. PMID 9407040.
^ Baldwin, Robert L. (June 1975). "Intermediates in Protein Folding Reactions and the Mechanism of Protein Folding". Annual Review of Biochemistry. 44 (1): 453–475. doi:10.1146/annurev.bi.44.070175.002321. ISSN 0066-4154. PMID 1094916.
^ Dabrowski-Tumanski, Pawel; Piejko, Maciej; Niewieczerzal, Szymon; Stasiak, Andrzej; Sulkowska, Joanna I. (2018-10-12). "Protein Knotting by Active Threading of Nascent Polypeptide Chain Exiting from the Ribosome Exit Channel". The Journal of Physical Chemistry B. 122 (49): 11616–11625. doi:10.1021/acs.jpcb.8b07634. ISSN 1520-6106. PMID 30198720.
^ Brandman O, et al. (2012). "A ribosome-bound quality control complex triggers degradation of nascent peptides and signals translation stress". Cell. 151 (5): 1042–54. doi:10.1016/j.cell.2012.10.044. PMC 3534965. PMID 23178123.
^ Defenouillère Q, et al. (2013). "Cdc48-associated complex bound to 60S particles is required for the clearance of aberrant translation products". Proc Natl Acad Sci U S A. 110 (13): 5046–51. doi:10.1073/pnas.1221724110. PMC 3612664. PMID 23479637.
^ Shen PS, et al. (2015). "Rqc2p and 60S ribosomal subunits mediate mRNA-independent elongation of nascent chains". Science. 347 (6217): 75–8. doi:10.1126/science.1259724. PMC 4451101. PMID 25554787.
^ Keeley, Jim; Gutnikoff, Robert (2015-01-02). "Ribosome Studies Turn Up New Mechanism of Protein Synthesis" (Press release). Howard Hughes Medical Institute. Archived from the original on 2015-01-12. Retrieved 2015-01-16.
^ University of Würzburg. (2016, October 6). Ribosomal quality control. ScienceDaily. "Ribosomal quality control". Archived from the original on 2017-11-07. Retrieved 2017-11-13.
^ http://newsrescue.com/ribosomal-quality-control/#axzz4qxPAgyHU Archived 2017-08-27 at the Wayback Machine
^ "Ribosomal quality control". Archived from the original on 2017-08-27. Retrieved 2017-08-27.
^ https://www.ncbi.nlm.nih.gov/books/NBK26841/#A2204 Archived 2017-10-03 at the Wayback Machine
^ ab Noller, H. F. (2012). "Evolution of protein synthesis from an RNA world". Cold Spring Harbor Perspectives in Biology. 4 (4): 1–U20. doi:10.1101/cshperspect.a003681. PMC 3312679. PMID 20610545.
^ Dabbs, E.R. (1986). Mutant studies on the prokaryotic ribosome. Springer-Verlag, N.Y.
^ Noller, H. F.; Hoffarth, V.; Zimniak, L. (June 5, 1992). "Unusual resistance of peptidyl transferase to protein extraction procedures". Science. 256 (5062): 1416–1419. doi:10.1126/science.1604315. PMID 1604315.
^ Nomura, M.; Mizushima, S.; Ozaki, M.; Trau, P.; Lowry, C. V. (1969). "Structure and function of ribosomes and their molecular components". Cold Spring Harbor Symposium of Quantitative Biology. 34: 49–61. doi:10.1101/sqb.1969.034.01.009.
^ ab Root-Bernstein, M.; Root-Bernstein, R. (2015). "The ribosome as a missing link in the evolution of life". Journal of Theoretical Biology. 367: 130–158. doi:10.1016/j.jtbi.2014.11.025. PMID 25500179.
^ Yarus M (2002). "Primordial genetics: phenotype of the ribocyte". Annu. Rev. Genet. 36: 125–51. doi:10.1146/annurev.genet.36.031902.105056. PMID 12429689.
^ Forterre, Patrick; Krupovic, Mart (2012). "The Origin of Virions and Virocells: The Escape Hypothesis Revisited". Viruses: Essential Agents of Life. pp. 43–60. doi:10.1007/978-94-007-4899-6_3. ISBN 978-94-007-4898-9.
^ Caetano-Anolles, G.; Seufferheld, M. J. (2013). "The coevolutionary roots of biochemistry and cellular organization challenge the RNA world paradigm". Journal of Molecular Microbiology and Biotechnology. 23 (1–2): 152–177. doi:10.1159/000346551. PMID 23615203.
^ Saladino, R.; Botta, G.; Pino, S.; Costanzo, G.; Mauro, E. Di (2012). "Genetics first or metabolism first? The formamide clue". Chemical Society Reviews. 41 (16): 5526–5565. doi:10.1039/c2cs35066a. PMID 22684046.
^ Bauer, Johann W.; Brandl, Clemens; Haubenreisser, Olaf; Wimmer, Bjoern; Weber, Manuela; Karl, Thomas; Klausegger, Alfred; Breitenbach, Michael; Hintner, Helmut (2013-07-08). "Specialized Yeast Ribosomes: A Customized Tool for Selective mRNA Translation". PLoS ONE. 8 (7): e67609. doi:10.1371/journal.pone.0067609. ISSN 1932-6203. PMC 3704640. PMID 23861776.
^ Mauro, Vincent P.; Edelman, Gerald M. (2002-09-17). "The ribosome filter hypothesis". Proceedings of the National Academy of Sciences of the United States of America. 99 (19): 12031–12036. doi:10.1073/pnas.192442499. ISSN 0027-8424. PMC 129393. PMID 12221294.
^ Xue, Shifeng; Barna, Maria (May 23, 2012). "Specialized ribosomes: a new frontier in gene regulation and organismal biology". Nature Reviews Molecular Cell Biology. 13 (6): 355–369. doi:10.1038/nrm3359. ISSN 1471-0080. PMC 4039366. PMID 22617470.
^ Mathis, Andrew (8 Dec 2016). "Mechanisms of in vivo ribosome maintenance change in response to nutrient signals". Molecular & Cellular Proteomics. mcp.M116.063255. (2): 243–254. doi:10.1074/mcp.M116.063255. PMC 5294211. PMID 27932527.
^ ab Guo, Huili (2018-08-20). "Specialized ribosomes and the control of translation". Biochemical Society Transactions. 46 (4): 855–869. doi:10.1042/BST20160426. ISSN 1470-8752. PMID 29986937.
^ Ballesta, J. P.; Guarinos, E.; Rodriguez-Gabriel, M. A.; Bermejo, B.; Jimenez-Diaz, A.; Remacha, M. (1995-09-01). "Ribosomal acidic phosphoproteins P1 and P2 are not required for cell viability but regulate the pattern of protein expression in Saccharomyces cerevisiae". Molecular and Cellular Biology. 15 (9): 4754–4762. doi:10.1128/MCB.15.9.4754. ISSN 1098-5549. PMC 230719. PMID 7651393.
^ Artero-Castro, Ana; Perez-Alea, Mileidys; Feliciano, Andrea; Leal, Jose A; Genestar, Mónica; Castellvi, Josep; Peg, Vicente; Ramón y Cajal, Santiago; LLeonart, Matilde E (2015-07-15). "Disruption of the ribosomal P complex leads to stress-induced autophagy". Autophagy. 11 (9): 1499–1519. doi:10.1080/15548627.2015.1063764. ISSN 1554-8627. PMC 4590587. PMID 26176264.
^ Marygold, S. J. (2005-02-01). "Genetic Analysis of RpL38 and RpL5, Two Minute Genes Located in the Centric Heterochromatin of Chromosome 2 of Drosophila melanogaster". Genetics. 169 (2): 683–695. doi:10.1534/genetics.104.034124. ISSN 0016-6731. PMC 1449105. PMID 15520262.
^ Kondrashov, Nadya; Pusic, Aya; Stumpf, Craig R.; Shimizu, Kunihiko; Hsieh, Andrew C.; Xue, Shifeng; Ishijima, Junko; Shiroishi, Toshihiko; Barna, Maria (April 2011). "Ribosome-Mediated Specificity in Hox mRNA Translation and Vertebrate Tissue Patterning". Cell. 145 (3): 383–397. doi:10.1016/j.cell.2011.03.028. ISSN 0092-8674. PMC 4445650. PMID 21529712.
^ Lee, S.-W.; Berger, S. J.; Martinovic, S.; Pasa-Tolic, L.; Anderson, G. A.; Shen, Y.; Zhao, R.; Smith, R. D. (2002-04-30). "Direct mass spectrometric analysis of intact proteins of the yeast large ribosomal subunit using capillary LC/FTICR". Proceedings of the National Academy of Sciences. 99 (9): 5942–5947. doi:10.1073/pnas.082119899. ISSN 0027-8424. PMC 122881. PMID 11983894.
^ Carroll, Adam J.; Heazlewood, Joshua L.; Ito, Jun; Millar, A. Harvey (2007-10-13). "Analysis of theArabidopsisCytosolic Ribosome Proteome Provides Detailed Insights into Its Components and Their Post-translational Modification". Molecular & Cellular Proteomics. 7 (2): 347–369. doi:10.1074/mcp.m700052-mcp200. ISSN 1535-9476. PMID 17934214.
^ Odintsova, Tatyana I.; Müller, Eva-Christina; Ivanov, Anton V.; Egorov, Tsezi A.; Bienert, Ralf; Vladimirov, Serguei N.; Kostka, Susanne; Otto, Albrecht; Wittmann-Liebold, Brigitte (2003). Journal of Protein Chemistry. 22 (3): 249–258. doi:10.1023/a:1025068419698. ISSN 0277-8033. Missing or empty|title=(help)
^ Yu, Yonghao; Ji, Hong; Doudna, Jennifer A.; Leary, Julie A. (2009-01-01). "Mass spectrometric analysis of the human 40S ribosomal subunit: Native and HCV IRES-bound complexes". Protein Science. 14 (6): 1438–1446. doi:10.1110/ps.041293005. ISSN 0961-8368. PMC 2253395. PMID 15883184.
^ Zeidan, Quira; Wang, Zihao; De Maio, Antonio; Hart, Gerald W. (2010-06-15). "O-GlcNAc Cycling Enzymes Associate with the Translational Machinery and Modify Core Ribosomal Proteins". Molecular Biology of the Cell. 21 (12): 1922–1936. doi:10.1091/mbc.e09-11-0941. ISSN 1059-1524. PMC 2883937. PMID 20410138.
^ Spence, Jean; Gali, Rayappa Reddy; Dittmar, Gunnar; Sherman, Fred; Karin, Michael; Finley, Daniel (July 2000). "Cell Cycle–Regulated Modification of the Ribosome by a Variant Multiubiquitin Chain". Cell. 102 (1): 67–76. doi:10.1016/s0092-8674(00)00011-8. ISSN 0092-8674. PMID 10929714.
^ Sauert, Martina; Temmel, Hannes; Moll, Isabella (July 2015). "Heterogeneity of the translational machinery: Variations on a common theme". Biochimie. 114: 39–47. doi:10.1016/j.biochi.2014.12.011. ISSN 0300-9084. PMC 4894553. PMID 25542647.
^ Thompson, Mary K; Rojas-Duran, Maria F; Gangaramani, Paritosh; Gilbert, Wendy V (2016-04-27). "The ribosomal protein Asc1/RACK1 is required for efficient translation of short mRNAs". eLife. 5. doi:10.7554/elife.11154. ISSN 2050-084X. PMC 4848094. PMID 27117520.
^ Landry, D. M.; Hertz, M. I.; Thompson, S. R. (2009-12-01). "RPS25 is essential for translation initiation by the Dicistroviridae and hepatitis C viral IRESs". Genes & Development. 23 (23): 2753–2764. doi:10.1101/gad.1832209. ISSN 0890-9369. PMC 2788332. PMID 19952110.
^ Deniz, N.; Lenarcic, E. M.; Landry, D. M.; Thompson, S. R. (2009-03-24). "Translation initiation factors are not required for Dicistroviridae IRES function in vivo". RNA. 15 (5): 932–946. doi:10.1261/rna.1315109. ISSN 1355-8382. PMC 2673076. PMID 19299549.
^ Hertz, M. I.; Landry, D. M.; Willis, A. E.; Luo, G.; Thompson, S. R. (2012-12-28). "Ribosomal Protein S25 Dependency Reveals a Common Mechanism for Diverse Internal Ribosome Entry Sites and Ribosome Shunting". Molecular and Cellular Biology. 33 (5): 1016–1026. doi:10.1128/mcb.00879-12. ISSN 0270-7306. PMC 3623076. PMID 23275440.
^ Decatur, Wayne A; Fournier, Maurille J (July 2002). "rRNA modifications and ribosome function". Trends in Biochemical Sciences. 27 (7): 344–351. doi:10.1016/s0968-0004(02)02109-6. ISSN 0968-0004.
^ Natchiar, S. Kundhavai; Myasnikov, Alexander G.; Kratzat, Hanna; Hazemann, Isabelle; Klaholz, Bruno P. (2017-11-15). "Visualization of chemical modifications in the human 80S ribosome structure". Nature. 551 (7681): 472–477. doi:10.1038/nature24482. ISSN 0028-0836. PMID 29143818.
External links
| Wikimedia Commons has media related to Ribosomes. |
- Lab computer simulates ribosome in motion
Role of the Ribosome, Gwen V. Childs, copied here
Ribosome in Proteopedia - The free, collaborative 3D encyclopedia of proteins & other molecules- Ribosomal proteins families in ExPASy
Molecule of the Month © RCSB Protein Data Bank:- Ribosome
- Elongation Factors
- Palade
- 3D electron microscopy structures of ribosomes at the EM Data Bank(EMDB)
![]() This article incorporates public domain material from the NCBI document "Science Primer".
This article incorporates public domain material from the NCBI document "Science Primer".